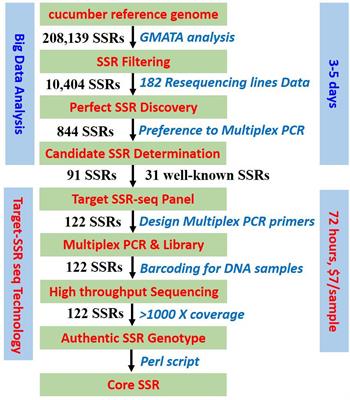
Frontiers | Target SSR-Seq: A Novel SSR Genotyping Technology Associate With Perfect SSRs in Genetic Analysis of Cucumber Varieties

SciELO - Brasil - Microsatellite markers: what they mean and why they are so useful Microsatellite markers: what they mean and why they are so useful
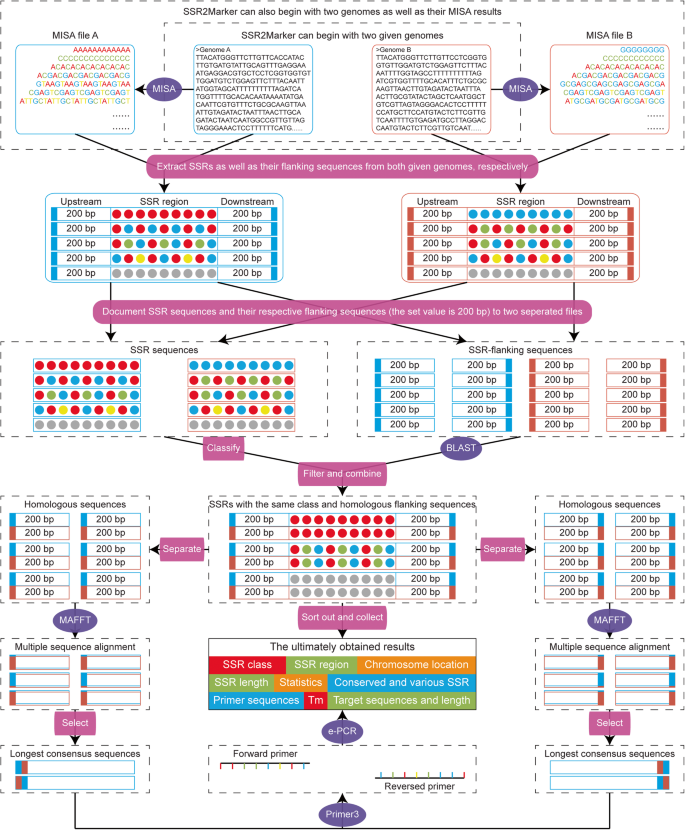
SSR2Marker: an integrated pipeline for identification of SSR markers within any two given genome-scale sequences | Molecular Horticulture | Full Text

Genes | Free Full-Text | Transferability and Polymorphism of SSR Markers Located in Flavonoid Pathway Genes in Fragaria and Rubus Species

Comprehensive functional analysis and mapping of SSR markers in the chickpea genome (Cicer arietinum L.) - ScienceDirect
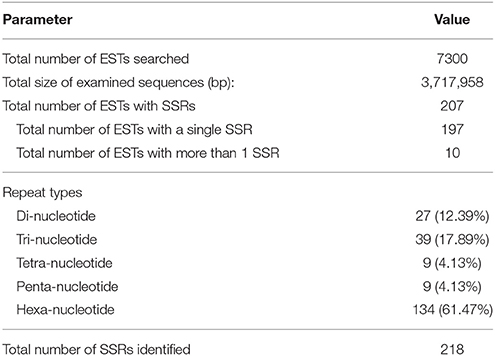
A Modified Method for the Development of SSR Molecular Markers Based on Redundant EST Data and Its Application in Soybean

Genome-wide simple sequence repeats (SSR) markers discovered from whole-genome sequence comparisons of multiple spinach accessions | Scientific Reports
Simple Sequence Repeats (SSR) Marker Screening Related to Orange Fleshed Sweet Potato F1 Genotype Resistance against Scab (Sphaceloma batatas Saw.) | KnE Life Sciences
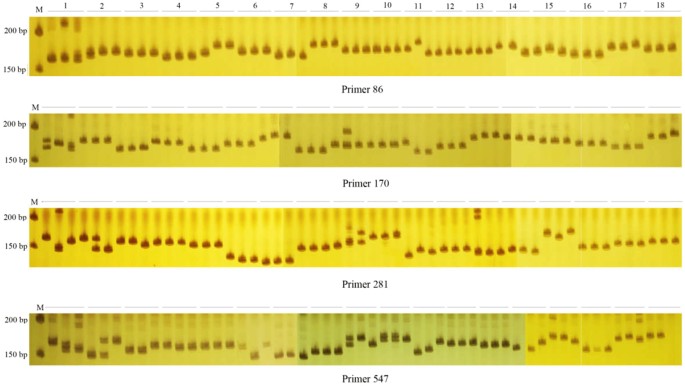

Cross-species transferability of EST-SSR markers developed from the transcriptome of Melilotus and their application to population genetics research | Scientific Reports

Genome-wide identification of simple sequence repeats and development of polymorphic SSR markers in swamp eel (Monopterus albus) - Hai-feng Tian, Qiao-mu Hu, Zhong Li, 2021
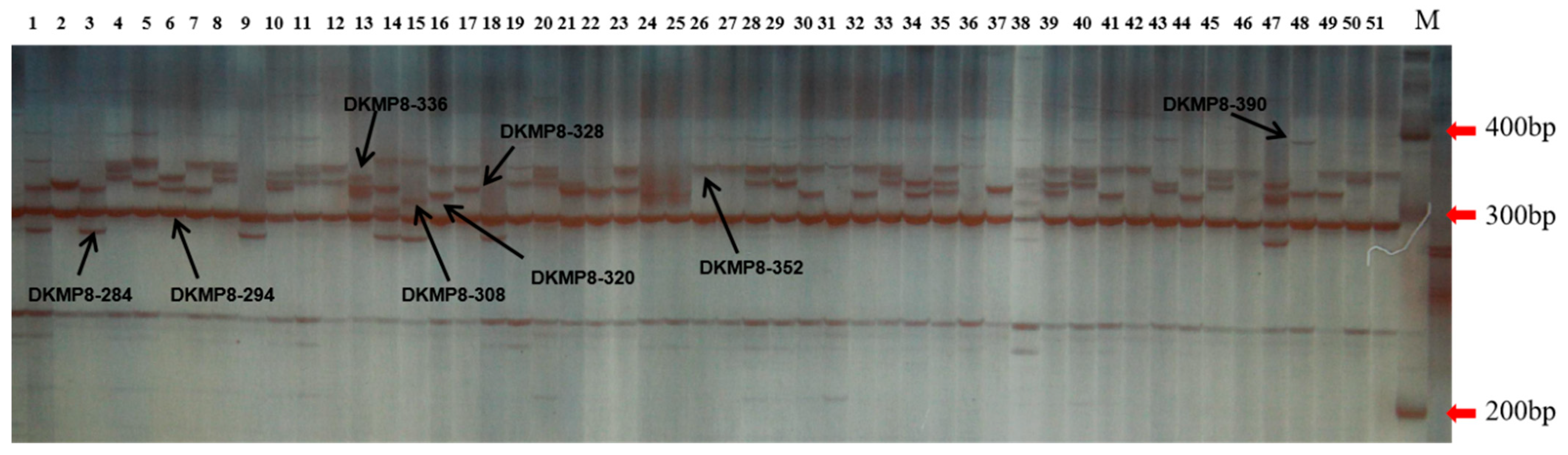
Forests | Free Full-Text | Genetic Diversity and Association Analysis among Germplasms of Diospyros kaki in Zhejiang Province Based on SSR Markers
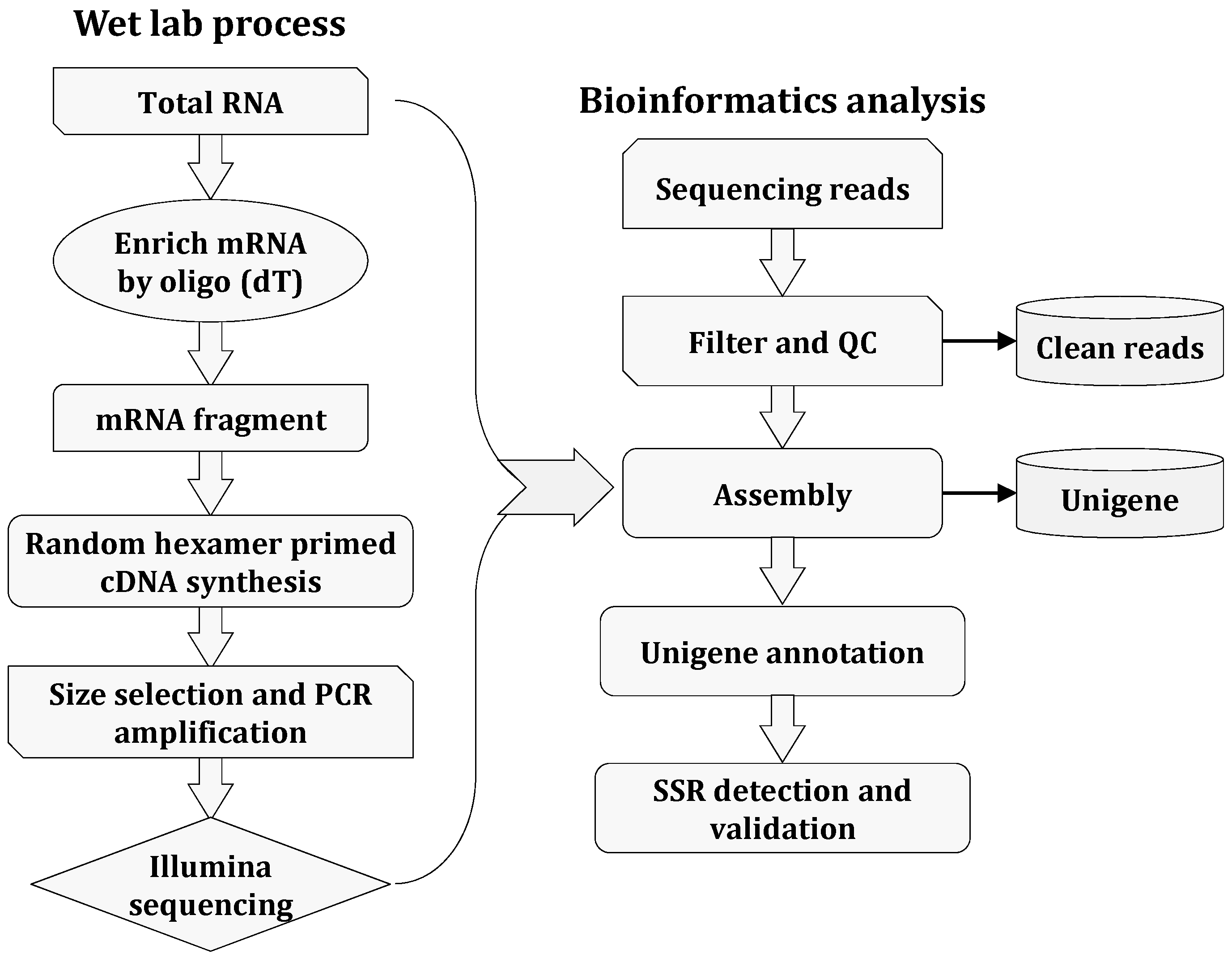
Molecules | Free Full-Text | Mining and Development of Novel SSR Markers Using Next Generation Sequencing (NGS) Data in Plants

Diversity | Free Full-Text | SSR Markers for Trichoderma virens: Their Evaluation and Application to Identify and Quantify Root-Endophytic Strains

Mining and validation of novel genotyping-by-sequencing (GBS)-based simple sequence repeats (SSRs) and their application for the estimation of the genetic diversity and population structure of coconuts (Cocos nucifera L.) in Thailand
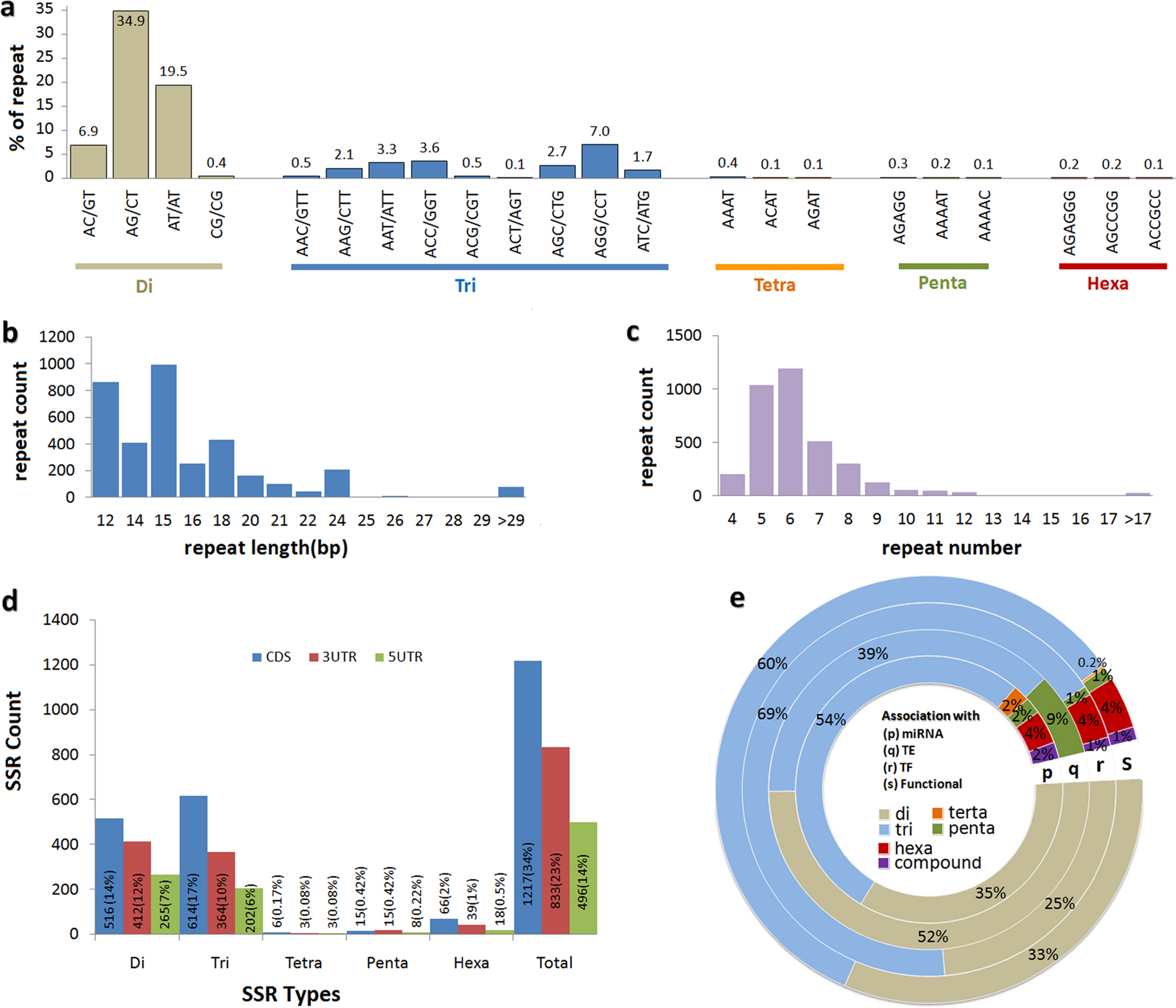
Transcriptome wide SSR discovery cross-taxa transferability and development of marker database for studying genetic diversity population structure of Lilium species | Scientific Reports
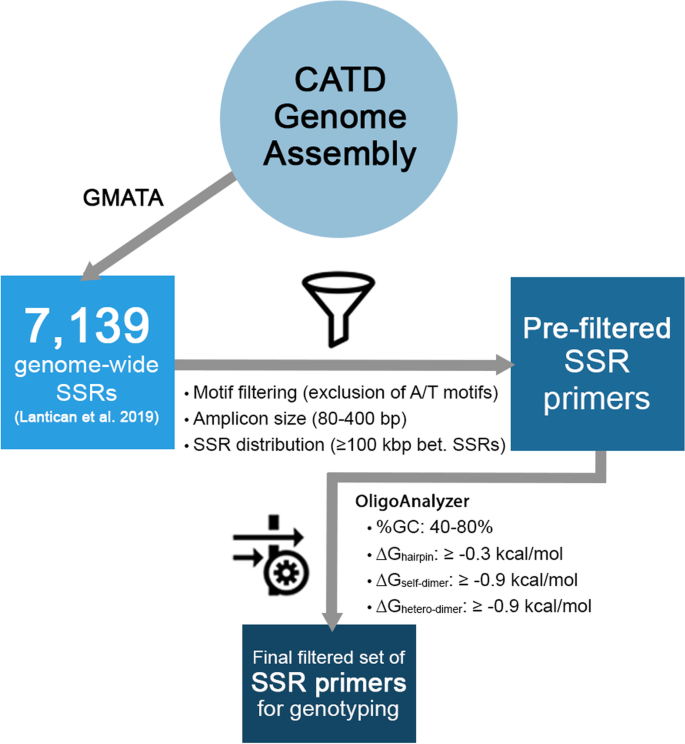
Mining and validation of novel simple sequence repeat (SSR) markers derived from coconut (Cocos nucifera L.) genome assembly | Journal of Genetic Engineering and Biotechnology | Full Text